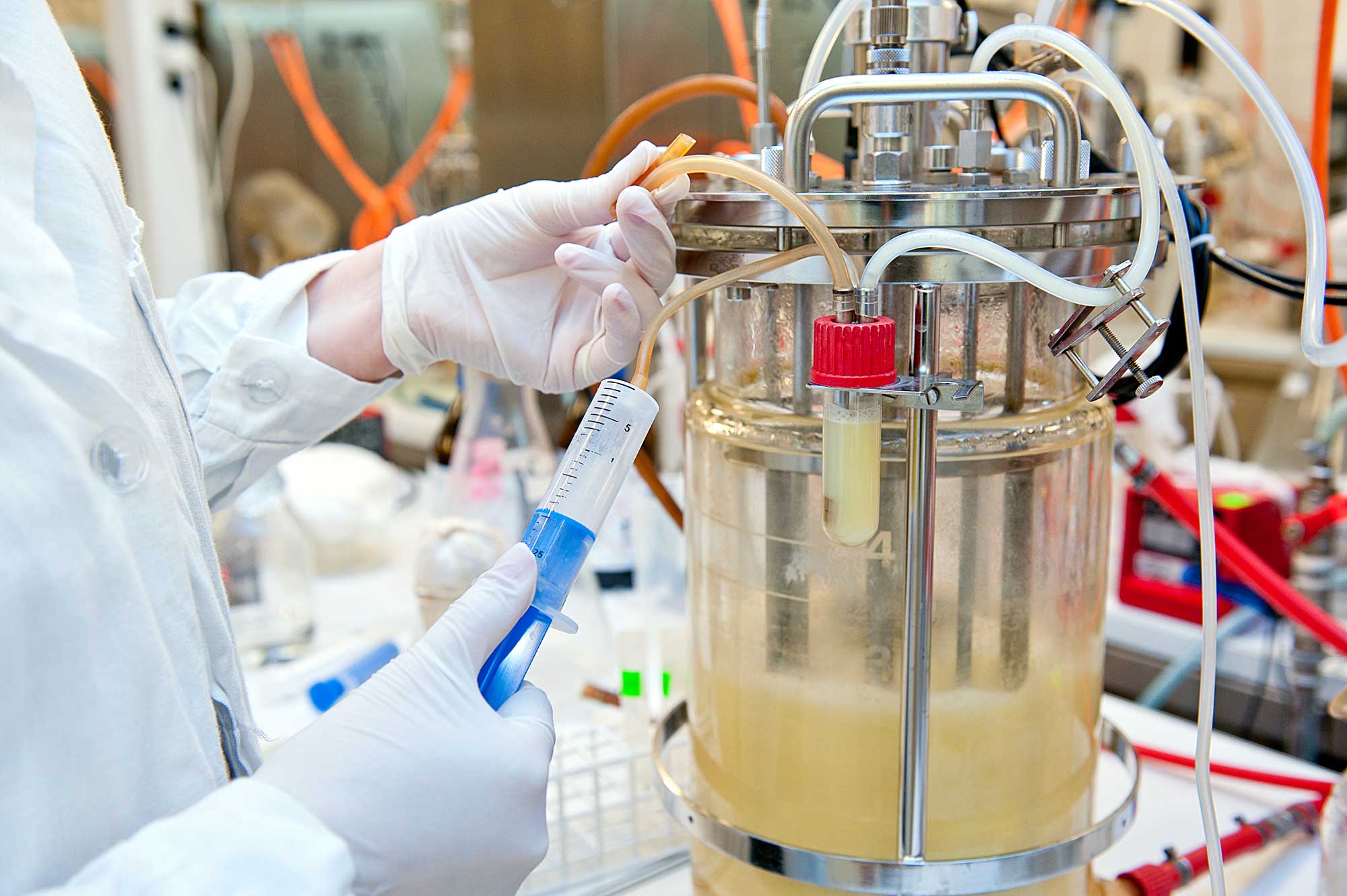

Theme I. Building Novel Microbial Manufacturing Platforms and Integrating Biocatalysis and Catalysis to Nurture a Sustainable Biorenewable Chemical Industry
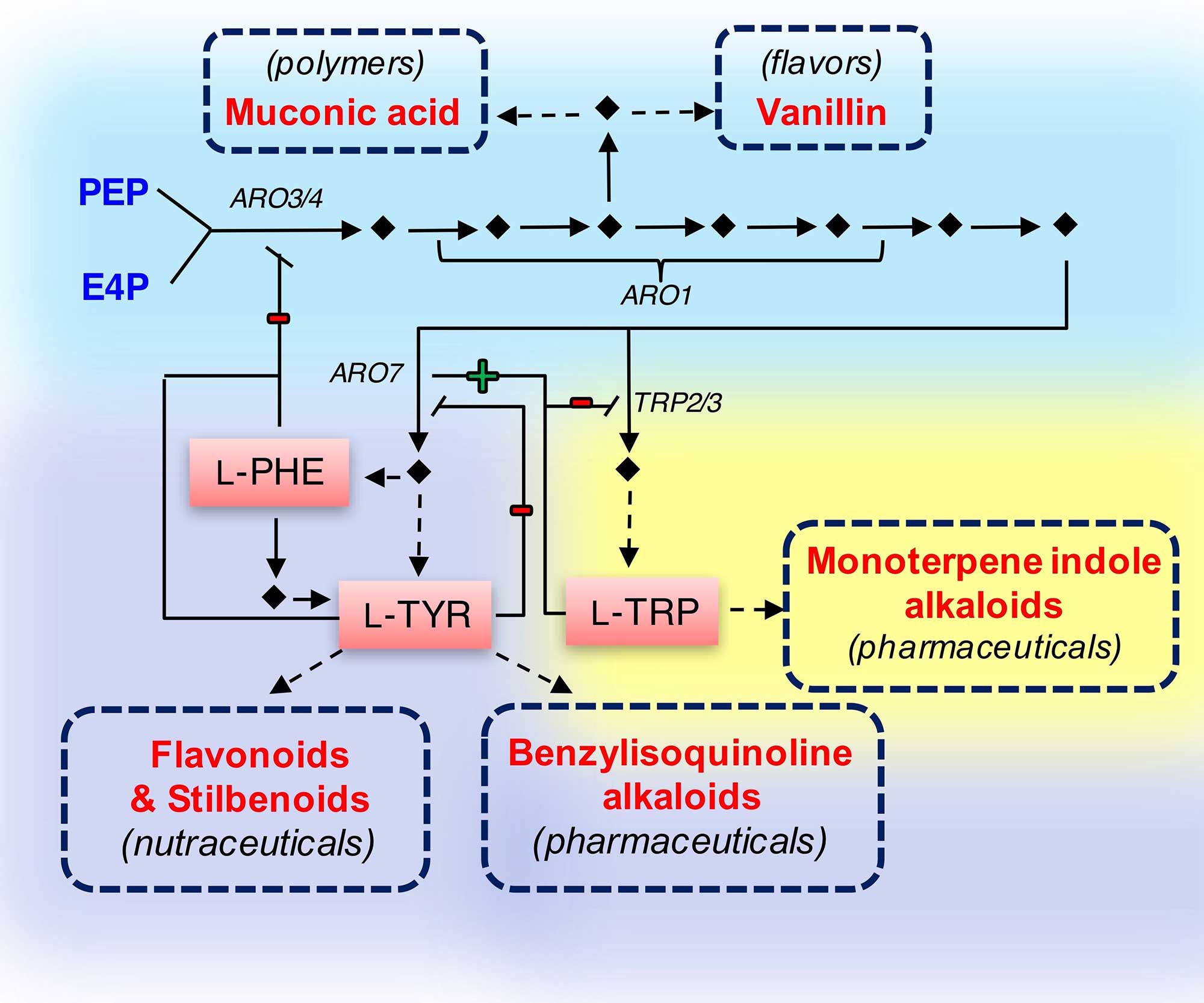
Our group designs various microbial factories for producing high-value compounds with applications in polymer precursors, nutraceuticals, and pharmaceuticals. Distinct from the mainstream research that is centered on exploiting the model microbial species, Our group emphasizes on “tailoring host selection according to the specific need.” For example, Scheffersomyces stipitis is explored to produce plant-sourced flavonoids and alkaloids (nutraceuticals and pharmaceuticals) because this microorganism renders higher precursor availability for the upstream shikimate pathway; Issatchenkia orientalis is being exploited to produce dicarboxylic acids (polymer precursors) owing to its superior acid tolerance; and Yarrowia lipolytica is selected as the producer to synthesize wax esters (premier lubricant and skin care additives) based on its exceptional oleaginous nature, which yields sufficient starting molecules.
In addition, aligning with the mission of the CBiRC to nurture a sustainable biorenewable chemical industry, our group has been collaborating with the chemists in CBiRC and polymer scientists to streamline the integration of biocatalysis and catalysis. For example, in association with ISU faculty Jean-Philippe Tessonnier and Eric Cochran, we developed a seamless integration platform to produce a nylon replacement molecule from biomass-derived sugar. At present, a collaborative team consisting of Tessonnier group, Broadbelt group (Northwestern University), and Shao group is striving to identify bioprivileged molecules for creating diverse chemical products including drop-in chemicals and novel species with desired functional replacements.
Theme II. Elucidating Core Design Principles to Streamline Nonconventional Microbial Host Development
In the course of selecting microbial production hosts, we recognized that the limited model microbial species are not the only hosts of potential scientific and economic importance. Numerous other nonconventional microorganisms exhibit highly unusual metabolic, biosynthetic, physiological, and fermentative capacities. As outcomes of long-term natural evolution in particular environments, these high-performance characteristics are conferred by a network of genes via a hierarchy of regulations that are intrinsically complex, rendering the horizontal transfer of these functions into model hosts highly challenging. However, significant technology hurdles lie in front of exploiting high potential nonconventional microorganisms.
This research theme is aimed at providing the main avenues for enabling rapid functional modifications of challenging nonconventional species and genome-scale genotype–phenotype profiling. At present, our group is establishing in silico principles to predict the genetic elements vital to the creation of stable expression platforms for convenient gene and pathway manipulations. We are also developing a high throughput pipeline for transcriptome engineering and reprogramming the genome-scale regulatory network to enhance the desired cellular function. This research is geared toward elucidating systematic rules and enabling generalizable platform technologies that can be effectively applied across species.


Theme III. Creating Microbial Consortia to Perform Complex Tasks
Microbial consortia, composed of multiple interacting microbial populations, can carry out complicated tasks that are more challenging for individual populations to perform. Cooperation and division of labor is common in nature, wherein organisms have established a wide variety of relationships. The co-culture technology has also been utilized in industrial processes such as food processing and waste degradation.
Our group is developing a yeast consortium to produce shikimate pathway derivatives with nutraceutical and pharmaceutical values. In a collaborative project with ISU faculty Laura Jarboe, we are also engineering an isogenic self-tuning bacterial consortium for converting mixed substrates and producing mixed products. In another collaborative project with ISU faculty Emily Smith and Ames Lab scientist Tanya Prozorov, our group is building microbial consortia and collaborating with the microscopy specialists to study the deconstruction of plant materials as an inexpensive feedstock for producing biofuels and chemicals.
Theme IV. Exploring Nucleosome-depleted Sequences for Novel Applications in Synthetic Biology
Functional genomics has been primarily focused on elucidating the roles of coding elements. Accumulating evidence demonstrates that the large amount of noncoding regions of a genome is not garbage. Their spatial arrangement and interaction with the coding elements play an important role in guiding cellular behaviors. In synthetic biology, the canonical Design–Build–Test–Learn cycle dissects complex natural biological subjects into distinct functional parts through a “subtraction” hierarchy. Numerous efforts are then focused on characterizing individual parts and modules at one layer of complexity at a time. However, the interplay among multiple layers in the new combinations interferes with the simplistic mix-and-match workflow and causes undesirable design imprecision.
Our group aims at addressing the challenge of the frequently encountered “low orthogonality” between well-characterized biological components and their genetic contexts using nucleosome-depleted sequences. We are seeking novel bioengineering solutions using these previously overlooked sequences to streamline the development of advanced microbial factories.